Machine learning generates 3D model from 2D pictures
Neural field network doesn’t need to be trained on other samples
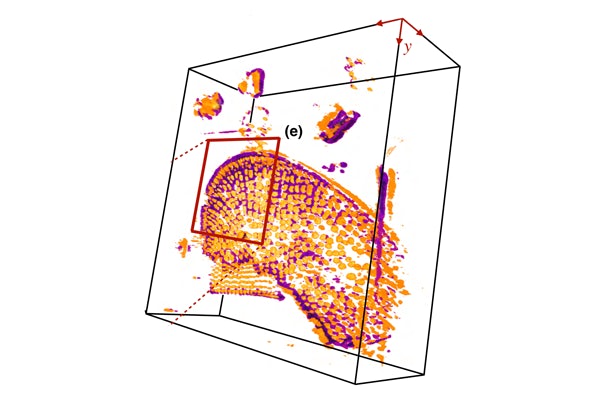
Researchers from the McKelvey School of Engineering at Washington University in St. Louis have developed a machine learning algorithm that can create a continuous 3D model of cells from a partial set of 2D images that were taken using the same standard microscopy tools found in many labs today.
Their findings were published Sept. 16 in the journal Nature Machine Intelligence.
“We train the model on the set of digital images to obtain a continuous representation,” said Ulugbek Kamilov, assistant professor of electrical and systems engineering and of computer science and engineering. “Now, I can show it any way I want. I can zoom in smoothly and there is no pixelation.”
The key to this work was the use of a neural field network, a particular kind of machine learning system that learns a mapping from spatial coordinates to the corresponding physical quantities. When the training is complete, researchers can point to any coordinate and the model can provide the image value at that location.
A particular strength of neural field networks is that they do not need to be trained on copious amounts of similar data. Instead, as long as there is a sufficient number of 2D images of the sample, the network can represent it in its entirety, inside and out.
The image used to train the network is just like any other microscopy image. In essence, a cell is lit from below; the light travels through it and is captured on the other side, creating an image.
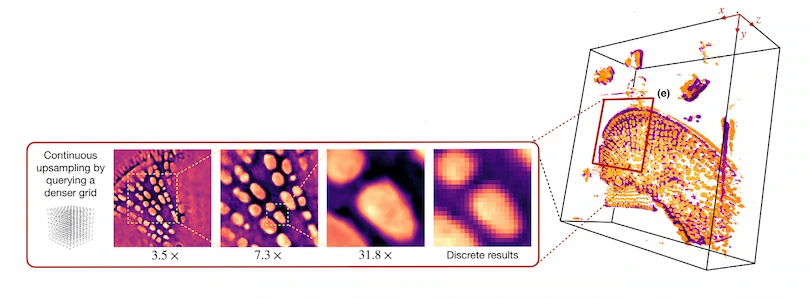
“Because I have some views of the cell, I can use those images to train the model,” Kamilov said. This is done by feeding the model information about a point in the sample where the image captured some of the internal structure of the cell.
Then the network takes its best shot at recreating that structure. If the output is wrong, the network is tweaked. If it’s correct, that pathway is reinforced. Once the predictions match real-world measurements, the network is ready to fill in parts of the cell that were not captured by the original 2D images.
The model now contains information of a full, continuous representation of the cell — there’s no need to save a data-heavy image file because it can always be recreated by the neural field network.
And, Kamilov said, not only is the model an easy-to-store, true representation of the cell, but also, in many ways, it’s more useful than the real thing.
“I can put any coordinate in and generate that view,” he said. “Or I can generate entirely new views from different angles.” He can use the model to spin a cell like a top or zoom in for a closer look; use the model to do other numerical tasks; or even feed it into another algorithm.
This work was supported by the National Science Foundation, awards CCF-1813910, CCF-2043134, CCF-1813848, and EPMD-1846784




