New structure found in cells
These ‘clusters’ have structures that encode function
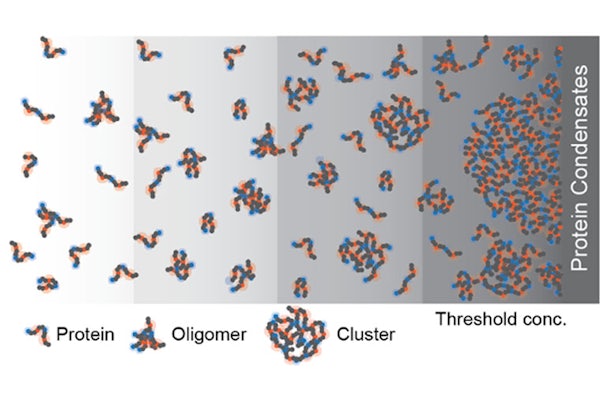
Every cell contains millions of protein molecules. Some of them have the ability to phase-separate to form non-membrane-bound compartments, called biomolecular condensates, inside a cell. It has long been assumed that there was no further structure underlying these condensates, only solution-soluble proteins.
A research group led by Rohit Pappu, the Gene K. Beare Distinguished Professor of biomedical engineering in the McKelvey School of Engineering at Washington University in St. Louis, and Anthony Hyman, director at the Max Planck Institute of Molecular Cell Biology and Genetics (MPI-CBG) in Dresden, Germany, uncovered that there is, in fact, relevant structure underlying condensates.
The finding, a collaboration with scientists at the University of Cambridge, Heinrich Heine University Düsseldorf and Technische Universität Dresden, was published July 5, 2022, in the Proceedings of the National Academy of Sciences.
The team found proteins formed dynamic structures at concentrations too low to form condensates. “Because of specific interactions, you can make lots of small ‘clusters,’” Pappu said. “The clusters have structures, and the structures encode function.”
Using dark-field microscopy, researchers tracked the motion of proteins in solution. They find clusters make up about 1% of the overall solution, even at protein concentrations just below the saturation level, where condensates form.
“Knowing that such clusters exist opens the door to assessing their functional relevance,” said Hyman, whose research group was responsible for initially discovering how widespread and relevant condensates are to cellular function.
The function of these protein clusters is still unknown and will be the subject of future studies. “Our findings highlight the totality of species that can form by proteins that are drivers of phase separation. Clearly, the next steps require that we determine the functions of clusters in subsaturated solutions because these concentrations at which they form are relevant in live cells,” Pappu said.
“Without a blueprint, we cannot describe the cellular process.”
Read more at the Max Planck Institute of Molecular Cell Biology and Genetics
This work was funded by a direct grant from the Max Planck Society, a grant from the NOMIS Foundation, the Wellcome Trust (No.209194/Z/17/Z ), the European Research Council (ERC) through ERC Grant PhysPro (aggreement337969), the Wellcome Trust and the Frances and Augustus Newman Foundation, priority program SPP2191 from the Deutsche Forschungsgemeinschaft, the US NIH (grants 5R01NS1056114 and R01NS121114), and the St. Jude Children’s Research Hospital collaborative research consortium on membraneless organelles.




