Distilling 70 years’ worth of data
Jonathan Silva and his lab use computational biology to better understand fundamentals of arrhythmia
In the heart, a complex set of proteins known as ion channels coordinates proper heartbeats. Dysregulation in the function of the ion channels leads to heart rhythm disorders, such as arrhythmia. Researchers in the McKelvey School of Engineering at Washington University in St. Louis recently sifted through hundreds of millions of models to find some that could help them get a better idea of how changes in the channel can destabilize the heart rhythm and lead to arrhythmia.
Jonathan Silva, associate professor of biomedical engineering, along with collaborators and members of his lab, used a method to identify all possible Markov models, which model the probabilities of different states, in ion channels. Ion channels can be either open or closed, representing two states. However, the channel goes through different states as its voltage, or action potential, changes across the cell membrane, which creates additional states.
In their initial computations, they found 111 million models possible for all of the various states. They eliminated duplicates and those that would be less useful for their work and ended up with 100,000 models. The team performed their computations using cloud computing resources from a grant from Amazon Web Services as well as in the McKelvey School of Engineering computing cluster. The method is described in PLoS Computational Biology Aug. 26, 2021. Kathryn Mangold, a graduate student in Silva’s lab, is first author of the paper.
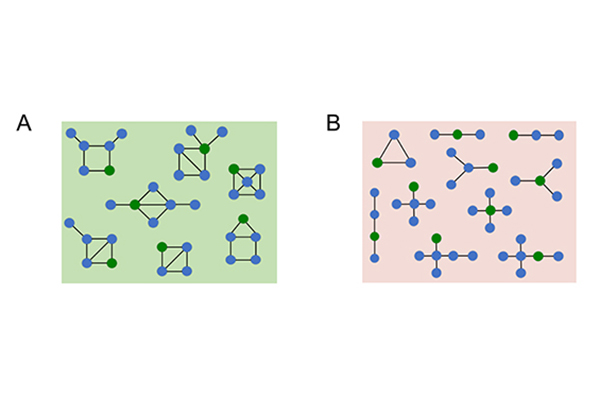
“We found models that you wouldn’t come up with intuitively because they connected in unusual ways,” Silva said. “That set of models gave us the idea of different ways to describe the channel behavior, and they were much simpler models, with five- or six-state models instead of 20-state models. It gave us an unbiased approach and allowed the data to guide the structure of the model.”
Once Silva’s team found the models, they wanted to connect the experimental data they observe when they characterize the ion channels to the way the channel contributes to the action potential and the heart muscle’s ability to contract.
“We do these very careful analyses of how the channels change at the molecular level, and sometimes it’s not clear how those changes will cause an arrhythmia,” Silva said.
The team put its computational channel models, which were based on data from the lab of Jeanne Nerbonne, professor of medicine and of developmental biology and director of the Center for Cardiovascular Research, both at the School of Medicine, into a mouse atrial myocyte model and into a human ventricular myocyte action potential model.
“We looked for reasons why the changes we see in an in vitro system cause an arrhythmia in a patient and why a drug that interacts with the channel in one way stabilizes the heart rhythm while other drugs are toxic,” Silva said.
Silva said this new framework helps them evaluate 70 years’ worth of ion channel data.
“When you have more and more results, it becomes difficult to connect it together,” he said. “By teaching the computer to do it, we can find the best model.”
Mangold K, Wang W, Johnson E, Bhagavan D, Moreno J, Nerbonne J, Silva J. Identification of Structures for Ion Channel Kinetic Models. PLoS Computational Biology, Aug. 26, 2021, https://doi.org/10.1371/journal.pcbi.1008932.
This work was supported by NIH grants R01HL136553, R01HL142520 and R01HL150637; the National Heart Lung and Blood sponsored institutional training grant (T32-HL134635); and Amazon Web Services Grant.




